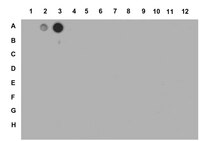
The Nucleosome Acidic Patch Regulates the H2B K123 Monoubiquitylation Cascade and Transcription Elongation in Saccharomyces cerevisiae.
Cucinotta, CE; Young, AN; Klucevsek, KM; Arndt, KM
PLoS genetics
11
e1005420
2015
Show Abstract
Eukaryotes regulate gene expression and other nuclear processes through the posttranslational modification of histones. In S. cerevisiae, the mono-ubiquitylation of histone H2B on lysine 123 (H2B K123ub) affects nucleosome stability, broadly influences gene expression and other DNA-templated processes, and is a prerequisite for additional conserved histone modifications that are associated with active transcription, namely the methylation of lysine residues in H3. While the enzymes that promote these chromatin marks are known, regions of the nucleosome required for the recruitment of these enzymes are undefined. To identify histone residues required for H2B K123ub, we exploited a functional interaction between the ubiquitin-protein ligase, Rkr1/Ltn1, and H2B K123ub in S. cerevisiae. Specifically, we performed a synthetic lethal screen with cells lacking RKR1 and a comprehensive library of H2A and H2B residue substitutions, and identified H2A residues that are required for H2B K123ub. Many of these residues map to the nucleosome acidic patch. The substitutions in the acidic patch confer varying histone modification defects downstream of H2B K123ub, indicating that this region contributes differentially to multiple histone modifications. Interestingly, substitutions in the acidic patch result in decreased recruitment of H2B K123ub machinery to active genes and defects in transcription elongation and termination. Together, our findings reveal a role for the nucleosome acidic patch in recruitment of histone modification machinery and maintenance of transcriptional integrity. | | | 26241481
 |
A comprehensive epigenome map of Plasmodium falciparum reveals unique mechanisms of transcriptional regulation and identifies H3K36me2 as a global mark of gene suppression.
Karmodiya, K; Pradhan, SJ; Joshi, B; Jangid, R; Reddy, PC; Galande, S
Epigenetics & chromatin
8
32
2015
Show Abstract
Role of epigenetic mechanisms towards regulation of the complex life cycle/pathogenesis of Plasmodium falciparum, the causative agent of malaria, has been poorly understood. To elucidate stage-specific epigenetic regulation, we performed genome-wide mapping of multiple histone modifications of P. falciparum. Further to understand the differences in transcription regulation in P. falciparum and its host, human, we compared their histone modification profiles.Our comprehensive comparative analysis suggests distinct mode of transcriptional regulation in malaria parasite by virtue of poised genes and differential histone modifications. Furthermore, analysis of histone modification profiles predicted 562 genes producing anti-sense RNAs and 335 genes having bidirectional promoter activity, which raises the intriguing possibility of RNA-mediated regulation of transcription in P. falciparum. Interestingly, we found that H3K36me2 acts as a global repressive mark and gene regulation is fine tuned by the ratio of activation marks to H3K36me2 in P. falciparum. This novel mechanism of gene regulation is supported by the fact that knockout of SET genes (responsible for H3K36 methylation) leads to up-regulation of genes with highest occupancy of H3K36me2 in wild-type P. falciparum. Moreover, virulence (var) genes are mostly poised and marked by a unique set of activation (H4ac) and repression (H3K9me3) marks, which are mutually exclusive to other Plasmodium housekeeping genes.Our study reveals unique plasticity in the epigenetic regulation in P. falciparum which can influence parasite virulence and pathogenicity. The observed differences in the histone code and transcriptional regulation in P. falciparum and its host will open new avenues for epigenetic drug development against malaria parasite. | | | 26388940
 |
Histone modifications rather than the novel regional centromeres of Zymoseptoria tritici distinguish core and accessory chromosomes.
Schotanus, K; Soyer, JL; Connolly, LR; Grandaubert, J; Happel, P; Smith, KM; Freitag, M; Stukenbrock, EH
Epigenetics & chromatin
8
41
2015
Show Abstract
Supernumerary chromosomes have been found in many organisms. In fungi, these "accessory" or "dispensable" chromosomes are present at different frequencies in populations and are usually characterized by higher repetitive DNA content and lower gene density when compared to the core chromosomes. In the reference strain of the wheat pathogen, Zymoseptoria tritici, eight discrete accessory chromosomes have been found. So far, no functional role has been assigned to these chromosomes; however, they have existed as separate entities in the karyotypes of Zymoseptoria species over evolutionary time. In this study, we addressed what-if anything-distinguishes the chromatin of accessory chromosomes from core chromosomes. We used chromatin immunoprecipitation combined with high-throughput sequencing ("ChIP-seq") of DNA associated with the centromere-specific histone H3, CENP-A (CenH3), to identify centromeric DNA, and ChIP-seq with antibodies against dimethylated H3K4, trimethylated H3K9 and trimethylated H3K27 to determine the relative distribution and proportion of euchromatin, obligate and facultative heterochromatin, respectively.Centromeres of the eight accessory chromosomes have the same sequence composition and structure as centromeres of the 13 core chromosomes and they are of similar length. Unlike those of most other fungi, Z. tritici centromeres are not composed entirely of repetitive DNA; some centromeres contain only unique DNA sequences, and bona fide expressed genes are located in regions enriched with CenH3. By fluorescence microscopy, we showed that centromeres of Z. tritici do not cluster into a single chromocenter during interphase. We found dramatically higher enrichment of H3K9me3 and H3K27me3 on the accessory chromosomes, consistent with the twofold higher proportion of repetitive DNA and poorly transcribed genes. In contrast, no single histone modification tested here correlated with the distribution of centromeric nucleosomes.All centromeres are similar in length and composed of a mixture of unique and repeat DNA, and most contain actively transcribed genes. Centromeres, subtelomeric regions or telomere repeat length cannot account for the differences in transfer fidelity between core and accessory chromosomes, but accessory chromosomes are greatly enriched in nucleosomes with H3K27 trimethylation. Genes on accessory chromosomes appear to be silenced by trimethylation of H3K9 and H3K27. | | | 26430472
 |
Lysine-specific demethylase (LSD1/KDM1A) and MYCN cooperatively repress tumor suppressor genes in neuroblastoma.
Amente, S; Milazzo, G; Sorrentino, MC; Ambrosio, S; Di Palo, G; Lania, L; Perini, G; Majello, B
Oncotarget
6
14572-83
2015
Show Abstract
The chromatin-modifying enzyme lysine-specific demethylase 1, KDM1A/LSD1 is involved in maintaining the undifferentiated, malignant phenotype of neuroblastoma cells and its overexpression correlated with aggressive disease, poor differentiation and infaust outcome. Here, we show that LSD1 physically binds MYCN both in vitro and in vivo and that such an interaction requires the MYCN BoxIII. We found that LSD1 co-localizes with MYCN on promoter regions of CDKN1A/p21 and Clusterin (CLU) suppressor genes and cooperates with MYCN to repress the expression of these genes. KDM1A needs to engage with MYCN in order to associate with the CDKN1A and CLU promoters. The expression of CLU and CDKN1A can be restored in MYCN-amplified cells by pharmacological inhibition of LSD1 activity or knockdown of its expression. Combined pharmacological inhibition of MYCN and LSD1 through the use of small molecule inhibitors synergistically reduces MYCN-amplified Neuroblastoma cell viability in vitro. These findings demonstrate that LSD1 is a critical co-factor of the MYCN repressive function, and suggest that combination of LSD1 and MYCN inhibitors may have strong therapeutic relevance to counteract MYCN-driven oncogenesis. | | | 26062444
 |
An integrative analysis of post-translational histone modifications in the marine diatom Phaeodactylum tricornutum.
Veluchamy, A; Rastogi, A; Lin, X; Lombard, B; Murik, O; Thomas, Y; Dingli, F; Rivarola, M; Ott, S; Liu, X; Sun, Y; Rabinowicz, PD; McCarthy, J; Allen, AE; Loew, D; Bowler, C; Tirichine, L
Genome biology
16
102
2015
Show Abstract
Nucleosomes are the building blocks of chromatin where gene regulation takes place. Chromatin landscapes have been profiled for several species, providing insights into the fundamental mechanisms of chromatin-mediated transcriptional regulation of gene expression. However, knowledge is missing for several major and deep-branching eukaryotic groups, such as the Stramenopiles, which include the diatoms. Diatoms are highly diverse and ubiquitous species of phytoplankton that play a key role in global biogeochemical cycles. Dissecting chromatin-mediated regulation of genes in diatoms will help understand the ecological success of these organisms in contemporary oceans.Here, we use high resolution mass spectrometry to identify a full repertoire of post-translational modifications on histones of the marine diatom Phaeodactylum tricornutum, including eight novel modifications. We map five histone marks coupled with expression data and show that P. tricornutum displays both unique and broadly conserved chromatin features, reflecting the chimeric nature of its genome. Combinatorial analysis of histone marks and DNA methylation demonstrates the presence of an epigenetic code defining activating or repressive chromatin states. We further profile three specific histone marks under conditions of nitrate depletion and show that the histone code is dynamic and targets specific sets of genes.This study is the first genome-wide characterization of the histone code from a stramenopile and a marine phytoplankton. The work represents an important initial step for understanding the evolutionary history of chromatin and how epigenetic modifications affect gene expression in response to environmental cues in marine environments. | | | 25990474
 |
An Lnc RNA (GAS5)/SnoRNA-derived piRNA induces activation of TRAIL gene by site-specifically recruiting MLL/COMPASS-like complexes.
He, X; Chen, X; Zhang, X; Duan, X; Pan, T; Hu, Q; Zhang, Y; Zhong, F; Liu, J; Zhang, H; Luo, J; Wu, K; Peng, G; Luo, H; Zhang, L; Li, X; Zhang, H
Nucleic Acids Res
43
3712-25
2015
Show Abstract
PIWI-interacting RNA (piRNA) silences the transposons in germlines or induces epigenetic modifications in the invertebrates. However, its function in the mammalian somatic cells remains unknown. Here we demonstrate that a piRNA derived from Growth Arrest Specific 5, a tumor-suppressive long non-coding RNA, potently upregulates the transcription of tumor necrosis factor (TNF)-related apoptosis-inducing ligand (TRAIL), a proapoptotic protein, by inducing H3K4 methylation/H3K27 demethylation. Interestingly, the PIWIL1/4 proteins, which bind with this piRNA, directly interact with WDR5, resulting in a site-specific recruitment of the hCOMPASS-like complexes containing at least MLL3 and UTX (KDM6A). We have indicated a novel pathway for piRNAs to specially activate gene expression. Given that MLL3 or UTX are frequently mutated in various tumors, the piRNA/MLL3/UTX complex mediates the induction of TRAIL, and consequently leads to the inhibition of tumor growth. | | | 25779046
 |
Distinct patterns of the histone marks associated with recruitment of the methionine chain-elongation pathway from leucine biosynthesis.
Xue, M; Long, J; Jiang, Q; Wang, M; Chen, S; Pang, Q; He, Y
Journal of experimental botany
66
805-12
2015
Show Abstract
Aliphatic glucosinolates (GLSs) are derived from chain-elongated methionine produced by an iterative three-step process, known to be evolutionarily recruited from leucine biosynthesis. The divergence of homologous genes between two pathways is mainly linked to the alterations in biochemical features. In this study, it was discovered that a distinct pattern of histone modifications is associated with and/or contributes to the divergence of the two pathways. In general, genes involved in leucine biosynthesis were robustly associated with H3k4me2 and H3K4me3. In contrast, despite the considerable abundances of H3K4me2 observed in some of genes involved in methionine chain elongation, H3K4me3 was completely missing. This H3K4m3-depleted pattern had no effect on gene transcription, whereas it seemingly co-evolved with the entire pathway of aliphatic GLS biosynthesis. The results reveal a novel association of the epigenetic marks with plant secondary metabolism, and may help to understand the recruitment of the methionine chain-elongation pathway from leucine biosynthesis. | | | 25428994
 |
Lysine-specific demethylase 2 suppresses lipid influx and metabolism in hepatic cells.
Nagaoka, K; Hino, S; Sakamoto, A; Anan, K; Takase, R; Umehara, T; Yokoyama, S; Sasaki, Y; Nakao, M
Molecular and cellular biology
35
1068-80
2015
Show Abstract
Cells link environmental fluctuations, such as nutrition, to metabolic remodeling. Epigenetic factors are thought to be involved in such cellular processes, but the molecular basis remains unclear. Here we report that the lysine-specific demethylase 2 (LSD2) suppresses the flux and metabolism of lipids to maintain the energy balance in hepatic cells. Using transcriptome and chromatin immunoprecipitation-sequencing analyses, we revealed that LSD2 represses the genes involved in lipid influx and metabolism through demethylation of histone H3K4. Selective recruitment of LSD2 at lipid metabolism gene loci was mediated in part by a stress-responsive transcription factor, c-Jun. Intriguingly, LSD2 depletion increased the intracellular levels of many lipid metabolites, which was accompanied by an increased susceptibility to toxic cell damage in response to fatty acid exposure. Our data demonstrate that LSD2 maintains metabolic plasticity under fluctuating environment in hepatocytes by mediating the cross talk between the epigenome and metabolism. | | | 25624347
 |
ROW1 maintains quiescent centre identity by confining WOX5 expression to specific cells.
Zhang, Y; Jiao, Y; Liu, Z; Zhu, YX
Nature communications
6
6003
2015
Show Abstract
The quiescent centre (QC) in the Arabidopsis root apical meristem is essential for stem cell organization. Here we show that the loss of REPRESSOR OF WUSCHEL1 (ROW1), a PHD domain-containing protein, leads to QC failure, defects in cell differentiation and ectopic expression of WUSCHEL-RELATED HOMEOBOX 5 (WOX5) in cells that normally express ROW1. The wox5-1/row1-3 double mutants show similar phenotypes to wox5-1 indicating that WOX5 is epistatic to ROW1. ROW1 specifically binds trimethylated histone H3 lysine 4 (H3K4me3) in the WOX5 promoter region to repress its transcription. QC expression of ROW1 results in a wox5-1-like phenotype with undetectable WOX5 transcripts. We propose that ROW1 is essential for QC maintenance and for stem cell niche development through the repression of WOX5 in the proximal meristem. | | | 25631790
 |
Poised chromatin and bivalent domains facilitate the mitosis-to-meiosis transition in the male germline.
Sin, HS; Kartashov, AV; Hasegawa, K; Barski, A; Namekawa, SH
BMC biology
13
53
2015
Show Abstract
The male germline transcriptome changes dramatically during the mitosis-to-meiosis transition to activate late spermatogenesis genes and to transiently suppress genes commonly expressed in somatic lineages and spermatogenesis progenitor cells, termed somatic/progenitor genes.These changes reflect epigenetic regulation. Induction of late spermatogenesis genes during spermatogenesis is facilitated by poised chromatin established in the stem cell phases of spermatogonia, whereas silencing of somatic/progenitor genes during meiosis and postmeiosis is associated with formation of bivalent domains which also allows the recovery of the somatic/progenitor program after fertilization. Importantly, during spermatogenesis mechanisms of epigenetic regulation on sex chromosomes are different from autosomes: X-linked somatic/progenitor genes are suppressed by meiotic sex chromosome inactivation without deposition of H3K27me3.Our results suggest that bivalent H3K27me3 and H3K4me2/3 domains are not limited to developmental promoters (which maintain bivalent domains that are silent throughout the reproductive cycle), but also underlie reversible silencing of somatic/progenitor genes during the mitosis-to-meiosis transition in late spermatogenesis. | | | 26198001
 |