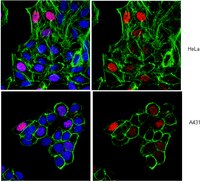
Regulation of p53 during senescence in normal human keratinocytes.
Kim, RH; Kang, MK; Kim, T; Yang, P; Bae, S; Williams, DW; Phung, S; Shin, KH; Hong, C; Park, NH
Aging cell
14
838-46
2015
Show Abstract
p53, the guardian of the genome, is a tumor suppressor protein and critical for the genomic integrity of the cells. Many studies have shown that intracellular level of p53 is enhanced during replicative senescence in normal fibroblasts, and the enhanced level of p53 is viewed as the cause of senescence. Here, we report that, unlike in normal fibroblasts, the level of intracellular p53 reduces during replicative senescence and oncogene-induced senescence (OIS) in normal human keratinocytes (NHKs). We found that the intracellular p53 level was also decreased in age-dependent manner in normal human epithelial tissues. Senescent NHKs exhibited an enhanced level of p16(INK4A) , induced G2 cell cycle arrest, and lowered the p53 expression and transactivation activity. We found that low level of p53 in senescent NHKs was due to reduced transcription of p53. The methylation status at the p53 promoter was not altered during senescence, but senescent NHKs exhibited notably lower level of acetylated histone 3 (H3) at the p53 promoter in comparison with rapidly proliferating cells. Moreover, p53 knockdown in rapidly proliferating NHKs resulted in the disruption of fidelity in repaired DNA. Taken together, our study demonstrates that p53 level is diminished during replicative senescence and OIS and that such diminution is associated with H3 deacetylation at the p53 promoter. The reduced intracellular p53 level in keratinocytes of the elderly could be a contributing factor for more frequent development of epithelial cancer in the elderly because of the loss of genomic integrity of cells. | | | 26138448
 |
Identification of in vivo DNA-binding mechanisms of Pax6 and reconstruction of Pax6-dependent gene regulatory networks during forebrain and lens development.
Sun, J; Rockowitz, S; Xie, Q; Ashery-Padan, R; Zheng, D; Cvekl, A
Nucleic acids research
43
6827-46
2015
Show Abstract
The transcription factor Pax6 is comprised of the paired domain (PD) and homeodomain (HD). In the developing forebrain, Pax6 is expressed in ventricular zone precursor cells and in specific subpopulations of neurons; absence of Pax6 results in disrupted cell proliferation and cell fate specification. Pax6 also regulates the entire lens developmental program. To reconstruct Pax6-dependent gene regulatory networks (GRNs), ChIP-seq studies were performed using forebrain and lens chromatin from mice. A total of 3514 (forebrain) and 3723 (lens) Pax6-containing peaks were identified, with ∼70% of them found in both tissues and thereafter called 'common' peaks. Analysis of Pax6-bound peaks identified motifs that closely resemble Pax6-PD, Pax6-PD/HD and Pax6-HD established binding sequences. Mapping of H3K4me1, H3K4me3, H3K27ac, H3K27me3 and RNA polymerase II revealed distinct types of tissue-specific enhancers bound by Pax6. Pax6 directly regulates cortical neurogenesis through activation (e.g. Dmrta1 and Ngn2) and repression (e.g. Ascl1, Fezf2, and Gsx2) of transcription factors. In lens, Pax6 directly regulates cell cycle exit via components of FGF (Fgfr2, Prox1 and Ccnd1) and Wnt (Dkk3, Wnt7a, Lrp6, Bcl9l, and Ccnd1) signaling pathways. Collectively, these studies provide genome-wide analysis of Pax6-dependent GRNs in lens and forebrain and establish novel roles of Pax6 in organogenesis. | | | 26138486
 |
Loss of EZH2 results in precocious mammary gland development and activation of STAT5-dependent genes.
Yoo, KH; Oh, S; Kang, K; Hensel, T; Robinson, GW; Hennighausen, L
Nucleic acids research
43
8774-89
2015
Show Abstract
Establishment and differentiation of mammary alveoli during pregnancy are controlled by prolactin through the transcription factors STAT5A and STAT5B (STAT5), which also regulate temporal activation of mammary signature genes. This study addressed the question whether the methyltransferase and transcriptional co-activator EZH2 controls the differentiation clock of mammary epithelium. Ablation of Ezh2 from mammary stem cells resulted in precocious differentiation of alveolar epithelium during pregnancy and the activation of mammary-specific STAT5 target genes. This coincided with enhanced occupancy of these loci by STAT5, EZH1 and RNA Pol II. Limited activation of differentiation-specific genes was observed in mammary epithelium lacking both EZH2 and STAT5, suggesting a modulating but not mandatory role for STAT5. Loss of EZH2 did not result in overt changes in genome-wide and gene-specific H3K27me3 profiles, suggesting compensation through enhanced EZH1 recruitment. Differentiated mammary epithelia did not form in the combined absence of EZH1 and EZH2. Transplantation experiments failed to demonstrate a role for EZH2 in the activity of mammary stem and progenitor cells. In summary, while EZH1 and EZH2 serve redundant functions in the establishment of H3K27me3 marks and the formation of mammary alveoli, the presence of EZH2 is required to control progressive differentiation of milk secreting epithelium during pregnancy. | | | 26250110
 |
A comprehensive epigenome map of Plasmodium falciparum reveals unique mechanisms of transcriptional regulation and identifies H3K36me2 as a global mark of gene suppression.
Karmodiya, K; Pradhan, SJ; Joshi, B; Jangid, R; Reddy, PC; Galande, S
Epigenetics & chromatin
8
32
2015
Show Abstract
Role of epigenetic mechanisms towards regulation of the complex life cycle/pathogenesis of Plasmodium falciparum, the causative agent of malaria, has been poorly understood. To elucidate stage-specific epigenetic regulation, we performed genome-wide mapping of multiple histone modifications of P. falciparum. Further to understand the differences in transcription regulation in P. falciparum and its host, human, we compared their histone modification profiles.Our comprehensive comparative analysis suggests distinct mode of transcriptional regulation in malaria parasite by virtue of poised genes and differential histone modifications. Furthermore, analysis of histone modification profiles predicted 562 genes producing anti-sense RNAs and 335 genes having bidirectional promoter activity, which raises the intriguing possibility of RNA-mediated regulation of transcription in P. falciparum. Interestingly, we found that H3K36me2 acts as a global repressive mark and gene regulation is fine tuned by the ratio of activation marks to H3K36me2 in P. falciparum. This novel mechanism of gene regulation is supported by the fact that knockout of SET genes (responsible for H3K36 methylation) leads to up-regulation of genes with highest occupancy of H3K36me2 in wild-type P. falciparum. Moreover, virulence (var) genes are mostly poised and marked by a unique set of activation (H4ac) and repression (H3K9me3) marks, which are mutually exclusive to other Plasmodium housekeeping genes.Our study reveals unique plasticity in the epigenetic regulation in P. falciparum which can influence parasite virulence and pathogenicity. The observed differences in the histone code and transcriptional regulation in P. falciparum and its host will open new avenues for epigenetic drug development against malaria parasite. | | | 26388940
 |
Impact of flanking chromosomal sequences on localization and silencing by the human non-coding RNA XIST.
Kelsey, AD; Yang, C; Leung, D; Minks, J; Dixon-McDougall, T; Baldry, SE; Bogutz, AB; Lefebvre, L; Brown, CJ
Genome biology
16
208
2015
Show Abstract
X-chromosome inactivation is a striking example of epigenetic silencing in which expression of the long non-coding RNA XIST initiates the heterochromatinization and silencing of one of the pair of X chromosomes in mammalian females. To understand how the RNA can establish silencing across millions of basepairs of DNA we have modelled the process by inducing expression of XIST from nine different locations in human HT1080 cells.Localization of XIST, depletion of Cot-1 RNA, perinuclear localization, and ubiquitination of H2A occurs at all sites examined, while recruitment of H3K9me3 was not observed. Recruitment of the heterochromatic features SMCHD1, macroH2A, H3K27me3, and H4K20me1 occurs independently of each other in an integration site-dependent manner. Silencing of flanking reporter genes occurs at all sites, but the spread of silencing to flanking endogenous human genes is variable in extent of silencing as well as extent of spread, with silencing able to skip regions. The spread of H3K27me3 and loss of H3K27ac correlates with the pre-existing levels of the modifications, and overall the extent of silencing correlates with the ability to recruit additional heterochromatic features.The non-coding RNA XIST functions as a cis-acting silencer when expressed from nine different locations throughout the genome. A hierarchy among the features of heterochromatin reveals the importance of interaction with the local chromatin neighborhood for optimal spread of silencing, as well as the independent yet cooperative nature of the establishment of heterochromatin by the non-coding XIST RNA. | | | 26429547
 |
Incorporation of histone H3.1 suppresses the lineage potential of skeletal muscle.
Harada, A; Maehara, K; Sato, Y; Konno, D; Tachibana, T; Kimura, H; Ohkawa, Y
Nucleic acids research
43
775-86
2015
Show Abstract
Lineage potential is triggered by lineage-specific transcription factors in association with changes in the chromatin structure. Histone H3.3 variant is thought to play an important role in the regulation of lineage-specific genes. To elucidate the function of H3.3 in myogenic differentiation, we forced the expression of GFP-H3.1 to alter the balance between H3.1 and H3.3 in mouse C2C12 cells that could be differentiated into myotubes. GFP-H3.1 replaced H3.3 in the regulatory regions of skeletal muscle (SKM) genes and induced a decrease of H3K4 trimethylation (H3K4me3) and increase of H3K27 trimethylation (H3K27me3). Similar results were obtained by H3.3 knockdown. In contrast, MyoD-dependent H3.3 incorporation into SKM genes in fibroblasts induced an increase of H3K4me3 and H3K27me3. In mouse embryos, a bivalent modification of H3K4me3 and H3K27me3 was formed on H3.3-incorporated SKM genes before embryonic skeletal muscle differentiation. These results suggest that lineage potential is established through a selective incorporation of specific H3 variants that governs the balance of histone modifications. | | | 25539924
 |
Coupling of T cell receptor specificity to natural killer T cell development by bivalent histone H3 methylation.
Dobenecker, MW; Kim, JK; Marcello, J; Fang, TC; Prinjha, R; Bosselut, R; Tarakhovsky, A
The Journal of experimental medicine
212
297-306
2015
Show Abstract
The fidelity of T cell immunity depends greatly on coupling T cell receptor signaling with specific T cell effector functions. Here, we describe a chromatin-based mechanism that enables integration of TCR specificity into definite T cell lineage commitment. Using natural killer T cells (iNKT cell) as a model of a T cell subset that differentiates in response to specific TCR signaling, we identified a key role of histone H3 lysine 27 trimethylation (H3K27me3) in coupling iNKT cell TCR specificity with the generation of iNKT cells. We found that the Zbtb16/PLZF gene promoter that drives iNKT cell differentiation possesses a bivalent chromatin state characterized by the simultaneous presence of negative and positive H3K27me3 and H3K4me3 modifications. Depletion of H3K27me3 at the Zbtb16/PLZF promoter leads to uncoupling of iNKT cell development from TCR specificity and is associated with accumulation of iNKT-like CD4(+) cells that express a non-iNKT cell specific T cell repertoire. In turn, stabilization of H3K27me3 leads to a drastic reduction of the iNKT cell population. Our data suggest that H3K27me3 levels at the bivalent Zbtb16/PLZF gene define a threshold enabling precise coupling of TCR specificity to lineage commitment. | | | 25687282
 |
Epigenetic Modifications of the PGC-1α Promoter during Exercise Induced Expression in Mice.
Lochmann, TL; Thomas, RR; Bennett, JP; Taylor, SM
PloS one
10
e0129647
2015
Show Abstract
The transcriptional coactivator, PGC-1α, is known for its role in mitochondrial biogenesis. Although originally thought to exist as a single protein isoform, recent studies have identified additional promoters which produce multiple mRNA transcripts. One of these promoters (promoter B), approximately 13.7 kb upstream of the canonical PGC-1α promoter (promoter A), yields alternative transcripts present at levels much lower than the canonical PGC-1α mRNA transcript. In skeletal muscle, exercise resulted in a substantial, rapid increase of mRNA of these alternative PGC-1α transcripts. Although the β2-adrenergic receptor was identified as a signaling pathway that activates transcription from PGC-1α promoter B, it is not yet known what molecular changes occur to facilitate PGC-1α promoter B activation following exercise. We sought to determine whether epigenetic modifications were involved in this exercise response in mouse skeletal muscle. We found that DNA hydroxymethylation correlated to increased basal mRNA levels from PGC-1α promoter A, but that DNA methylation appeared to play no role in the exercise-induced activation of PGC-1α promoter B. The level of the activating histone mark H3K4me3 increased with exercise 2-4 fold across PGC-1α promoter B, but remained unaltered past the canonical PGC-1α transcriptional start site. Together, these data show that epigenetic modifications partially explain exercise-induced changes in the skeletal muscle mRNA levels of PGC-1α isoforms. | | | 26053857
 |
Lysine-specific demethylase (LSD1/KDM1A) and MYCN cooperatively repress tumor suppressor genes in neuroblastoma.
Amente, S; Milazzo, G; Sorrentino, MC; Ambrosio, S; Di Palo, G; Lania, L; Perini, G; Majello, B
Oncotarget
6
14572-83
2015
Show Abstract
The chromatin-modifying enzyme lysine-specific demethylase 1, KDM1A/LSD1 is involved in maintaining the undifferentiated, malignant phenotype of neuroblastoma cells and its overexpression correlated with aggressive disease, poor differentiation and infaust outcome. Here, we show that LSD1 physically binds MYCN both in vitro and in vivo and that such an interaction requires the MYCN BoxIII. We found that LSD1 co-localizes with MYCN on promoter regions of CDKN1A/p21 and Clusterin (CLU) suppressor genes and cooperates with MYCN to repress the expression of these genes. KDM1A needs to engage with MYCN in order to associate with the CDKN1A and CLU promoters. The expression of CLU and CDKN1A can be restored in MYCN-amplified cells by pharmacological inhibition of LSD1 activity or knockdown of its expression. Combined pharmacological inhibition of MYCN and LSD1 through the use of small molecule inhibitors synergistically reduces MYCN-amplified Neuroblastoma cell viability in vitro. These findings demonstrate that LSD1 is a critical co-factor of the MYCN repressive function, and suggest that combination of LSD1 and MYCN inhibitors may have strong therapeutic relevance to counteract MYCN-driven oncogenesis. | | | 26062444
 |
miR-142-5p and miR-130a-3p are regulated by IL-4 and IL-13 and control profibrogenic macrophage program.
Su, S; Zhao, Q; He, C; Huang, D; Liu, J; Chen, F; Chen, J; Liao, JY; Cui, X; Zeng, Y; Yao, H; Su, F; Liu, Q; Jiang, S; Song, E
Nature communications
6
8523
2015
Show Abstract
Macrophages play a pivotal role in tissue fibrogenesis, which underlies the pathogenesis of many end-stage chronic inflammatory diseases. MicroRNAs are key regulators of immune cell functions, but their roles in macrophage's fibrogenesis have not been characterized. Here we show that IL-4 and IL-13 induce miR-142-5p and downregulate miR-130a-3p in macrophages; these changes sustain the profibrogenic effect of macrophages. In vitro, miR-142-5p mimic prolongs STAT6 phosphorylation by targeting its negative regulator, SOCS1. Blocking miR-130a relieves its inhibition of PPARγ, which coordinates STAT6 signalling. In vivo, inhibiting miR-142-5p and increasing miR-130a-3p expression with locked nucleic acid-modified oligonucleotides inhibits CCL4-induced liver fibrosis and bleomycin-induced lung fibrosis in mice. Furthermore, macrophages from the tissue samples of patients with liver cirrhosis and idiopathic pulmonary fibrosis display increased miR-142-5p and decreased miR-130a-3p expression. Therefore, miR-142-5p and miR-130a-3p regulate macrophage profibrogenic gene expression in chronic inflammation. | | | 26436920
 |


























[221189-ALL].jpg)